Recently, Carl Zimmer made a criticism of the computer animations of molecular events (it’s the same criticism I made 8 years ago): they’re beautiful and they’re informative, but they leave out the critical aspect of stochastic behavior that is important in understanding the biochemistry. He’s talking specifically about kinesin, a transport protein which the animators are particularly fond of illustrating.
Every now and then, a tiny molecule loaded with fuel binds to one of the kinesin “feet.” It delivers a jolt of energy, causing that foot to leap off the molecular cable and flail wildly, pulling hard on the foot that’s still anchored. Eventually, the gyrating foot stumbles into contact again with the cable, locking on once more — and advancing the vesicle a tiny step forward. This updated movie offers a better way to picture our most intricate inner workings…. In the 2006 version, we can’t help seeing intention in the smooth movements of the molecules; it’s as if they’re trying to get from one place to another. In reality, however, the parts of our cells don’t operate with the precise movements of the springs and gears of a clock. They flail blindly in the crowd.
The illusion of directed, purposeful movement is a simplifying shortcut: as Zimmer describes, there actually is a lot of noise in the system, it’s just that the thermodynamics of the interactions promote a directionality to the motion. This is Chemistry 101. I figured that everyone with an undergraduate level of understanding of molecules would be able to grasp this.
I did not take into account willful ignorance, however. Jonathan Wells is angry that anyone dared to question the perfect “stately grace” of molecular machines, and accuses proponents of stochastic motion of Flailing Blindly: The Pseudoscience of Josh Rosenau and Carl Zimmer. He has a Ph.D. in biology, and he doesn’t understand what I just said was Chem 101? For shame.
But that’s not what the biological evidence shows. In fact, kinesin moves quickly, with precise movements, to get from one place to another. A kinesin molecule takes one 8-nanometer “step” along a microtubule for every high-energy ATP molecule it uses, and it uses about 80 ATPs per second. On the scale of a living cell, this movement is very fast. To visualize it on a macroscopic scale, imagine a microtubule as a one-lane road and the kinesin molecule as an automobile. The kinesin would be traveling over 200 miles per hour!
The speed of the reaction doesn’t say anything about the specifics of the molecular movement…and it’s especially not convincing when your trick is to multiply the actual speed in the cell by approximately 1012 to scale it up to the size of a car. The flow rate of the Mississippi river is about 1.5 miles per hour here in Minnesota — if you multiply that by 1012, oh my god, the water is moving at about 2000 times the speed of light!
But let’s set aside the stupid inflation for a minute. Wells cites a couple of papers to back up his claim of the rate of ATP consumption. It’s true. But it doesn’t show that the movement is steady and machine-like and precise at all. He must be trusting us to not bother even reading the paper.
Here’s the deal: we can actually watch single molecules of kinesin behaving. The typical trick is to use a fluorescent bead, attach that to kinesin, and then record the glowing bead’s movement as it is moving along with optical-trapping interferometry. That’s the problem with Wells’ accusation: we actually see the behavior, and it’s not linear, smooth, and graceful.
This is the data that the paper used to measure the quantum, jerky behavior of kinesin. Just look at the top graph: that’s a record of the bead’s movement over time. You should be able to see that the line holds steady at one distance for variable lengths of time, and then jerks upward. The “jerks” are distances of about 8nm, and the other graphs are power spectra to show that there is a peak periodicity of 8nm. It shows the opposite of what Wells claims; there are long pauses and sudden shifts in the directly observed track of kinesin movement. The 8nm emerges because when one “foot” of kinesin releases and wobbles forward to connect to tubulin, it has an 8nm step.
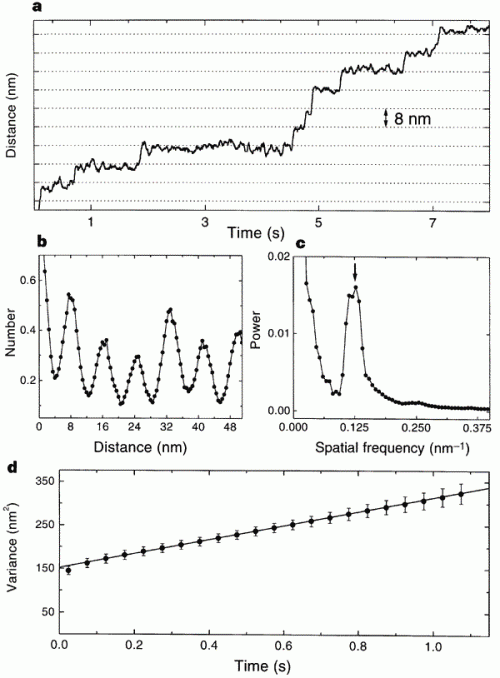
Horizontal grid lines (dotted lines) are spaced 8 nm apart. Data were median-filtered with a window width of 60 ms. b, Normalized histogram of pairwise distances between all pairs of data points in this record, showing a clear 8-nm periodicity. c, Normalized power spectrum of the data in b, displaying a single prominent peak at the reciprocal of 8 nm (arrow). d, Variance in position, averaged over 28 runs at 2 microM ATP (dots), and line fit over the interval 3.5 ms to 1.1 s. The y-intercept of the fit is determined by equipartition, left fencex2right fence = kT/alpha, where alpha is the combined stiffness of the optical trap and bead–microtubule linkage. The rapid rise in variance at short times reflects the brownian correlation time for bead position.
You can find lots of papers with direct observations of kinesin movement. Here’s data from another paper that essentially shows that the two ‘feet’ of kinesin alternate in their movement, because they made recombinant, asymmetric kinesin with slightly different step distances. Again, note the long dwell times punctuated with surges of movement.
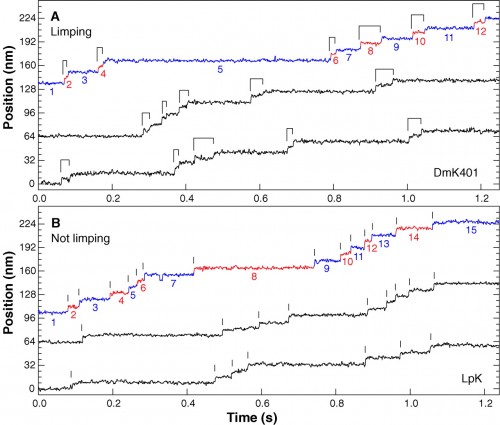
Representative high-resolution stepping records of position against time, showing the single-molecule behavior of kinesin motors under constant 4 pN rearward loads. (A) Limping motion of the recombinant kinesin construct, DmK401. The dwell intervals between successive 8-nm steps alternate between slow and fast phases, causing steps to appear in pairs, as indicated by the ligatures. (B) Nonlimping motion of native squid kinesin, LpK. No alternation of steps is apparent; vertical lines mark the stepping transitions. Slow and fast phase assignments, as described in the text, are indicated in color on the uppermost trace of each panel (blue and red, respectively), and the corresponding dwell intervals are numbered. All traces were median filtered with a 2.5-ms window.
The contrast is between “stately grace” and “jiggle and jump”. The evidence shows that it is the latter. Yet, somehow, Wells closes his weird series of non sequiturs with this question, as if he expects everyone to give him the answer he wants.
So, who are the pseudoscientists?
The answer is obvious. Wells and his cronies at the DI.
Asbury CL, Fehr AN, Block SM (2003) Kinesin moves by an asymmetric hand-over-hand mechanism. Science 302(5653):2130-4
Schnitzer MJ, Block SM (1997) Kinesin hydrolyses one ATP per 8-nm step. Nature388(6640):386-90.
By the way, I found about this because the Discovery Institute sent me email (silly season and all) proudly announcing that they were about to release evidence that would clearly rebut Zimmer’s assertion that kinesin “flails blindly” — they are making a computer animation showing that it doesn’t.
Yes, they are that stupid.