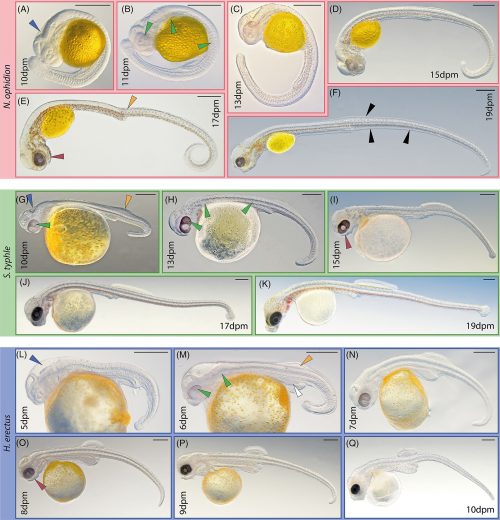
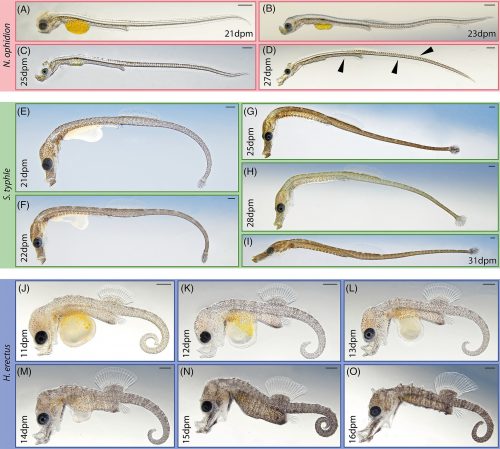
Show me a 500 million year old chelicerate, and I’ll be happy for a day. Look at this beauty, Megachelicerax cousteaui, excavated from a Utah fossil bed.
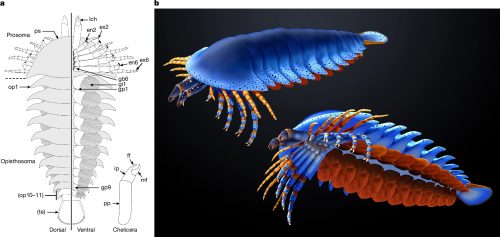
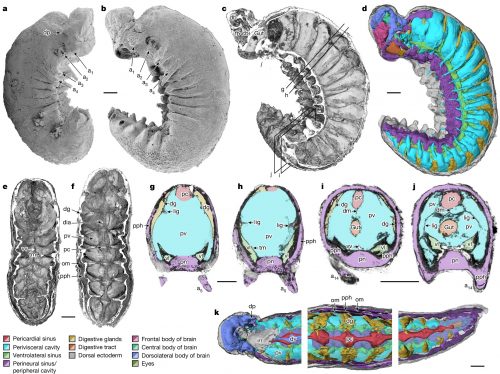
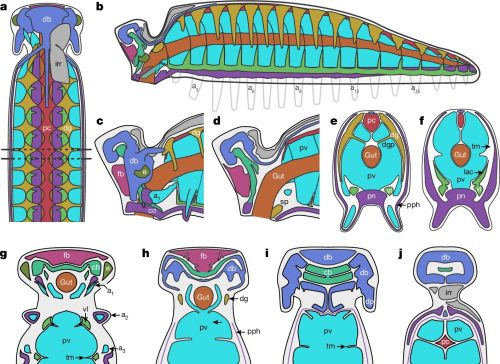
Anatomic reconstructions of the dorsal (left) and ventral (right) morphologies. b, Artistic reconstructions by M. Hattori illustrating oblique views of the dorsal (top) and ventral (bottom) morphologies. The sanctacaridid-like morphology of the posteriormost body region is speculative. gi, gill (that is, a set of gill lamellae); te, telson.
Pretty cool, right? The best part of it is that pair of appendages at the very front of the animal — those are chelicerae, the biting/chomping/chewing/venom-injecting bits of a modern spider, that make them distinct from insects, which only have antennae at that end. That makes this the oldest known chelicerate ever discovered. It was a swimming marine animal, and doesn’t have the legs we associate with spiders — chelicerae evolved first, legs much later.
Also, this isn’t just the mother of spiders, but is also the mother of a huge family of cousins: horseshoe crabs, eurypterids, as well as spiders.
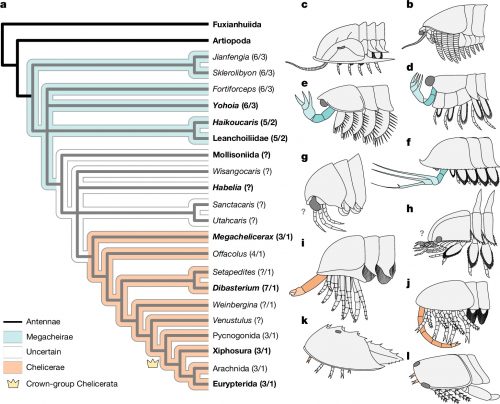
Megachelicerax documents the oldest stratigraphic occurrence of chelicerae (that is, uniramous, unichelate deutocerebral appendages) and bridges the simple body and limb organization of Cambrian megacheirans with the more derived anatomy of post-Cambrian synziphosurines and crown-group chelicerates. a, Simplified consensus topology based on Bayesian analysis (Mk model, 4 chains, 5,000,000 generations, 1/1,000 sampling resulting in 5,000 samples with 25% burn-in resulting in 3,750 samples retained); detailed results and comparison with parsimony provided in Extended Data Fig. 6. The numbers in parentheses correspond to the total number of podomeres and the number of chelae, respectively, present in the deutocerebral appendage. Taxa whose names are in bold font are illustrated in b–l. b–l, The morphology of the anterior body region in select taxa. b, Fuxianhuiid Chengjiangocaris kunmingensis (Cambrian, Stage 3). c, Artiopod Olenoides serratus (Cambrian, Wuliuan). d, Megacheiran Yohoia tenuis (Cambrian, Wuliuan). e, Megacheiran Haikoucaris ercaensis (Cambrian, Stage 3). f, Megacheiran Leanchoilia superlata (Cambrian, Wuliuan). g, Mollisoniid M. plenovenatrix (Cambrian, Wuliuan). h, Habeliid Habelia optata (Cambrian, Wuliuan). i, M. cousteaui (Cambrian, Drumian). j, Synziphosurine Dibasterium durgae (Silurian, Wenlock). k, Xiphosurid Limulus polyphemus (recent). l, Eurypterid Slimonia acuminata (Silurian, Llandovery–Wenlock).
That is one wildly successful tree. It just goes to show that you can go on to do great things even if your face looks like a nest of spiky clawed jointed tentacles.
Rudy Lerosey-Aubril, Javier Ortega-Hernández. A chelicera-bearing arthropod reveals the Cambrian origin of chelicerates. Nature, 2026; DOI: 10.1038/s41586-026-10284-2